---
title: "Classification — Logistic Regression and Metrics"
execute:
enabled: true
jupyter: python3
---
In Acts 1 and 2, we built and tested models for continuous outcomes. Act 3 extends these ideas to new settings, starting with classification — predicting categories instead of numbers. You used `RandomForestClassifier` back in Chapter 6. Today we understand *why* classification needs its own tools — and build a more interpretable classifier.
## The 85% accuracy trap
A hospital wants to predict which patients will be readmitted within 30 days, so they can intervene early. Missing those patients costs hospitals \$25,000+ per avoidable readmission in CMS penalties. But there's a catch: only 15% of patients are readmitted. If you just predict "not readmitted" for everyone, you're right 85% of the time.
**Is that a good model?** An 85%-accurate model that catches zero at-risk patients is worse than useless — it's expensive.
Today we'll see why accuracy is one of the most misleading metrics in statistics — and what to use instead.
```{python}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
sns.set_style('whitegrid')
plt.rcParams['figure.figsize'] = (7, 4.5)
plt.rcParams['font.size'] = 12
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import (confusion_matrix, ConfusionMatrixDisplay,
roc_curve, auc, precision_recall_curve)
from sklearn.preprocessing import StandardScaler
# Load data
DATA_DIR = 'data'
```
## The Framingham Heart Study
:::{.callout-note}
## The Framingham Heart Study
The Framingham Heart Study has tracked cardiovascular disease risk factors since 1948, following three generations of participants in Framingham, Massachusetts. It transformed our understanding of risk factors like cholesterol and blood pressure.
:::
Our dataset has 4,240 patients with 16 columns (15 predictors plus the outcome): did they develop coronary heart disease (CHD) within 10 years?
Let's look at the data and the class balance.
```{python}
# Load Framingham data
framingham = pd.read_csv(f'{DATA_DIR}/framingham/framingham.csv')
print(f"Shape: {framingham.shape[0]:,} rows x {framingham.shape[1]} columns")
framingham.head()
```
Now let's check the class balance: how many patients actually developed CHD?
```{python}
# Class balance: how many patients developed CHD?
chd_counts = framingham['TenYearCHD'].value_counts()
print("10-year CHD outcomes:")
print(f" No CHD: {chd_counts[0]:,} ({chd_counts[0]/len(framingham)*100:.1f}%)")
print(f" CHD: {chd_counts[1]:,} ({chd_counts[1]/len(framingham)*100:.1f}%)")
print(f" Ratio: {chd_counts[0]/chd_counts[1]:.1f}:1")
```
Only 15.2% of patients developed CHD. This ratio reflects **class imbalance** — the positive class is much rarer than the negative class. Class imbalance arises frequently in real applications: fraud detection, disease screening, equipment failure.
Class imbalance is going to cause us a lot of trouble today. Let's see why.
```{python}
# Prepare features and target
# Drop rows with missing values (we'll discuss imputation in other lectures)
framingham_clean = framingham.dropna()
print(f"After dropping NaN: {len(framingham_clean):,} rows (dropped {len(framingham) - len(framingham_clean):,})")
features = ['male', 'age', 'cigsPerDay', 'totChol', 'sysBP', 'diaBP',
'BMI', 'heartRate', 'glucose']
X = framingham_clean[features]
y = framingham_clean['TenYearCHD']
# Train/test split
# We use stratify=y to ensure the train and test sets have the same positive rate.
# Without this, random chance could give us a test set with very few CHD cases.
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=42, stratify=y
)
print(f"Training: {len(X_train):,} | Test: {len(X_test):,}")
print(f"Test positive rate: {y_test.mean():.3f}")
```
Let's look at the feature scales before we do anything else. Notice how different the ranges are — age is in years, blood pressure in mmHg, BMI around 20-40. This variation will matter when we interpret coefficients.
```{python}
# Feature scales -- notice the wide variation
X_train.describe().round(1)
```
## From regression to classification
In linear regression, we predicted a continuous outcome: price, blood pressure, etc. Now we want to predict a **binary** outcome: CHD or no CHD.
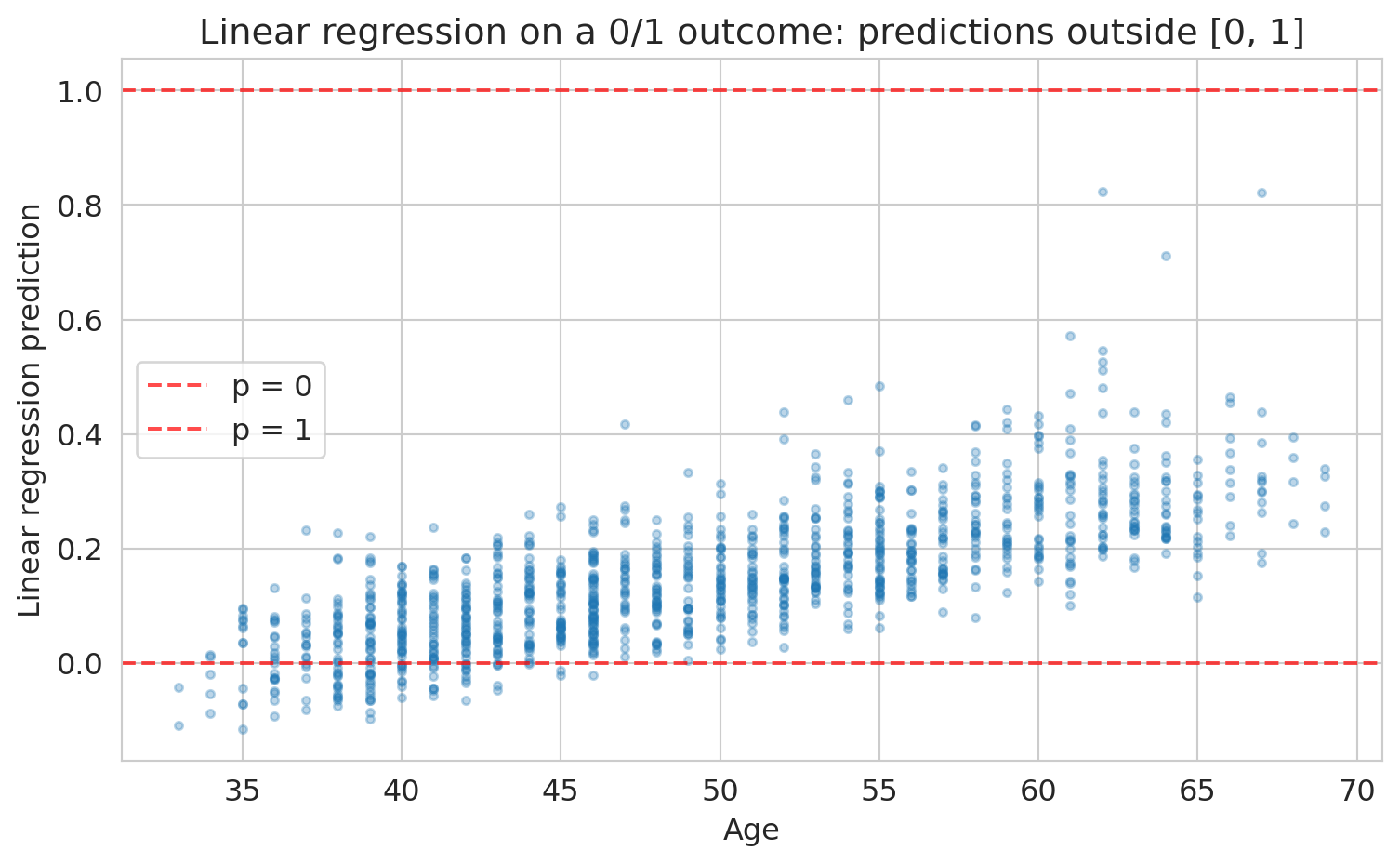
Could we just use linear regression on the 0/1 outcome? Let's try it and see what goes wrong.
```{python}
# What happens if we use linear regression on a binary outcome?
from sklearn.linear_model import LinearRegression
scaler_temp = StandardScaler()
X_train_temp = scaler_temp.fit_transform(X_train)
lin_model = LinearRegression().fit(X_train_temp, y_train)
lin_preds = lin_model.predict(scaler_temp.transform(X_test))
fig, ax = plt.subplots(figsize=(8, 5))
ax.scatter(X_test['age'], lin_preds, alpha=0.3, s=10)
ax.axhline(y=0, color='red', linestyle='--', alpha=0.7, label='p = 0')
ax.axhline(y=1, color='red', linestyle='--', alpha=0.7, label='p = 1')
ax.set_xlabel('Age')
ax.set_ylabel('Linear regression prediction')
ax.set_title('Linear regression on a 0/1 outcome: predictions outside [0, 1]')
ax.legend()
plt.tight_layout()
plt.show()
print(f"Predictions below 0: {(lin_preds < 0).sum()}")
print(f"Predictions above 1: {(lin_preds > 1).sum()}")
```
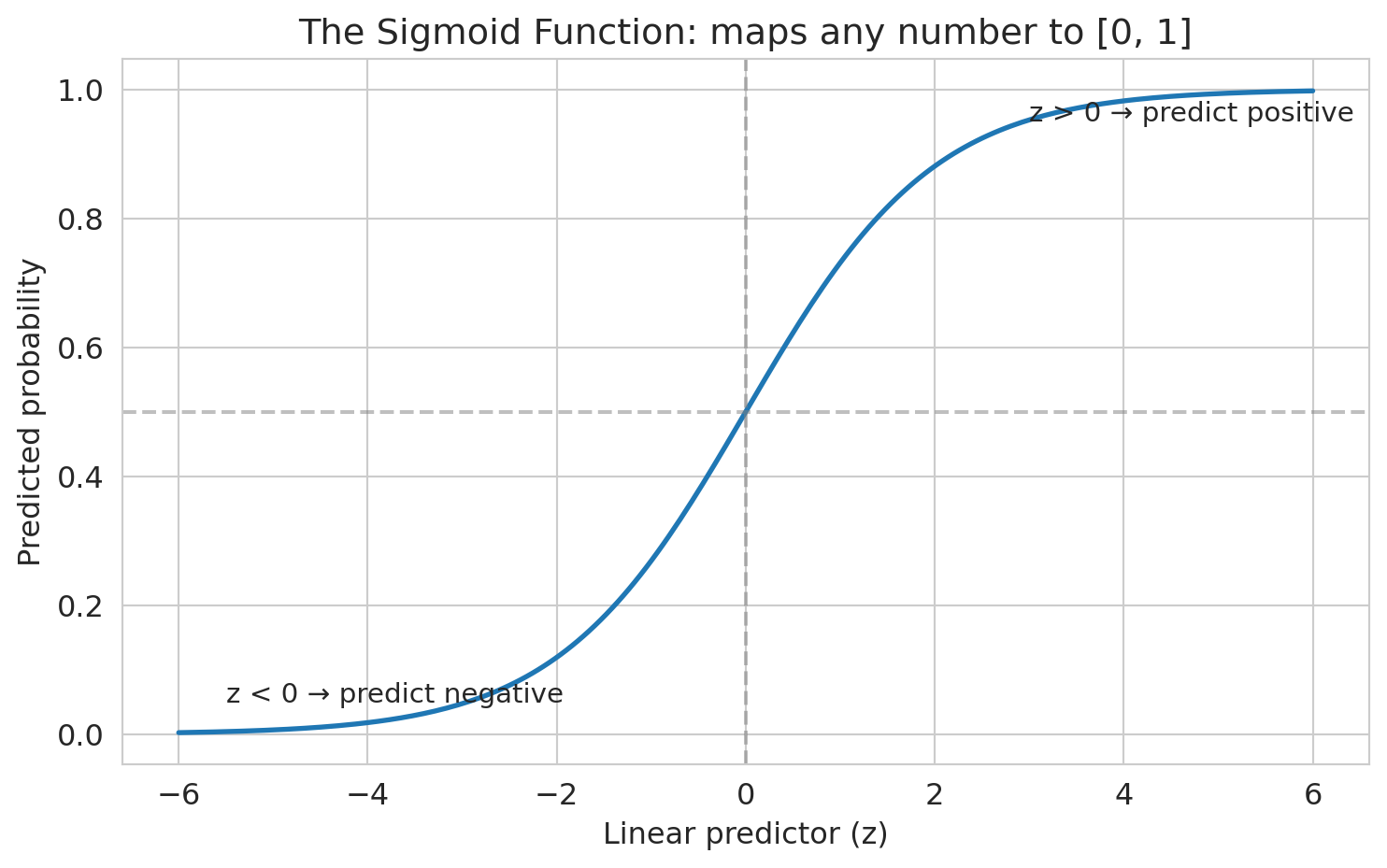
Some predictions are negative, some are above 1. Those don't make sense as probabilities. We need a function that squashes any number into [0, 1]. The **sigmoid function** does exactly this.
:::{.callout-important}
## Definition: Logistic Regression
**Logistic regression** predicts the **probability** that the outcome is 1:
$$p = \frac{1}{1 + e^{-(\beta_0 + \beta_1 x_1 + \cdots + \beta_p x_p)}}$$
The linear combination $\beta_0 + \beta_1 x_1 + \cdots$ can range from $-\infty$ to $+\infty$, but the sigmoid squashes it into a valid probability.
Equivalently, the **log-odds** (**logit**) is a linear function of the features:
$$\log\!\left(\frac{p}{1-p}\right) = \beta_0 + \beta_1 x_1 + \cdots + \beta_p x_p$$
:::
A concrete example makes the log-odds scale tangible. If $p = 0.20$: odds $= 0.20/0.80 = 0.25$, log-odds $= \log(0.25) = -1.39$. If $p = 0.80$: odds $= 0.80/0.20 = 4$, log-odds $= \log(4) = 1.39$. The symmetry is no accident — the log-odds scale is centered at zero (where $p = 0.5$) and stretches symmetrically toward $\pm\infty$.
:::{.callout-warning}
## Logistic Regression Conditions
There are two key conditions for logistic regression to be valid: (1) each outcome is independent of the others, and (2) each predictor is linearly related to the log-odds when all other predictors are held constant. Because the log-odds are linear in the features, the decision boundary (where $p = 0.5$) is a hyperplane — logistic regression can only produce linear decision boundaries.
:::
The name *logistic* regression reflects this core assumption: the log-odds are a **linear** function of the features. If the true decision boundary is curved, logistic regression will miss it — add polynomial or interaction features (Chapter 6) to capture nonlinear boundaries.
```{python}
# Visualize the sigmoid function
z = np.linspace(-6, 6, 200)
sigmoid = 1 / (1 + np.exp(-z))
fig, ax = plt.subplots(figsize=(8, 5))
ax.plot(z, sigmoid, linewidth=2)
ax.axhline(y=0.5, color='gray', linestyle='--', alpha=0.5)
ax.axvline(x=0, color='gray', linestyle='--', alpha=0.5)
ax.set_xlabel('Linear predictor (z)')
ax.set_ylabel('Predicted probability')
ax.set_title('The Sigmoid Function: maps any number to [0, 1]')
ax.annotate('z > 0 → predict positive', xy=(3, 0.95), fontsize=11)
ax.annotate('z < 0 → predict negative', xy=(-5.5, 0.05), fontsize=11)
plt.tight_layout()
plt.show()
```
## What does logistic regression optimize?
Why not just use squared error on the 0/1 outcomes? You could, but squared error doesn't penalize confident wrong predictions enough. A prediction of 0.9 for a true 0 has squared error $(0.9)^2 = 0.81$ — but the logistic loss for that same mistake is $-\log(0.1) = 2.3$, much larger. The worse the mistake, the more logistic loss amplifies the penalty.
> *"The loss function determines what the model finds."*
Logistic regression minimizes the **logistic loss** (also called **cross-entropy** or negative log-likelihood). The idea: we choose coefficients to maximize the likelihood of the observed outcomes. If a patient had CHD and we predicted probability 0.9, that's good (high likelihood). If we predicted 0.1 for a patient who had CHD, that's bad (low likelihood). Mathematically, the logistic loss for a single observation is:
$$\ell(\beta) = -\left[ y \log(p) + (1-y) \log(1-p) \right]$$
This loss penalizes confident wrong predictions *much* more heavily than squared error does. The model is punished harshly for saying "1% chance" when the patient actually develops CHD.
## Fitting logistic regression: gradient descent
In Chapter 5, we solved linear regression with the normal equations — a closed-form solution. Logistic regression has no such shortcut: the sigmoid makes the loss nonlinear. Instead, we use **gradient descent**, an iterative algorithm that repeatedly steps in the direction that decreases the loss most steeply.
The key formula: $\beta \leftarrow \beta - \eta \cdot \nabla L(\beta)$ where $\eta$ is the **learning rate** (step size). The logistic loss is **convex** — one valley, one global minimum — so gradient descent finds THE answer regardless of starting point.
```{python}
# Standardize features
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
```
Now let's fit the final model using sklearn's `LogisticRegression`, which uses a sophisticated version of this same idea under the hood.
```{python}
# Fit logistic regression
model = LogisticRegression(penalty=None, max_iter=1000, random_state=42)
model.fit(X_train_scaled, y_train)
print("Logistic regression fitted.")
print(f"Training accuracy: {model.score(X_train_scaled, y_train):.3f}")
print(f"Test accuracy: {model.score(X_test_scaled, y_test):.3f}")
```
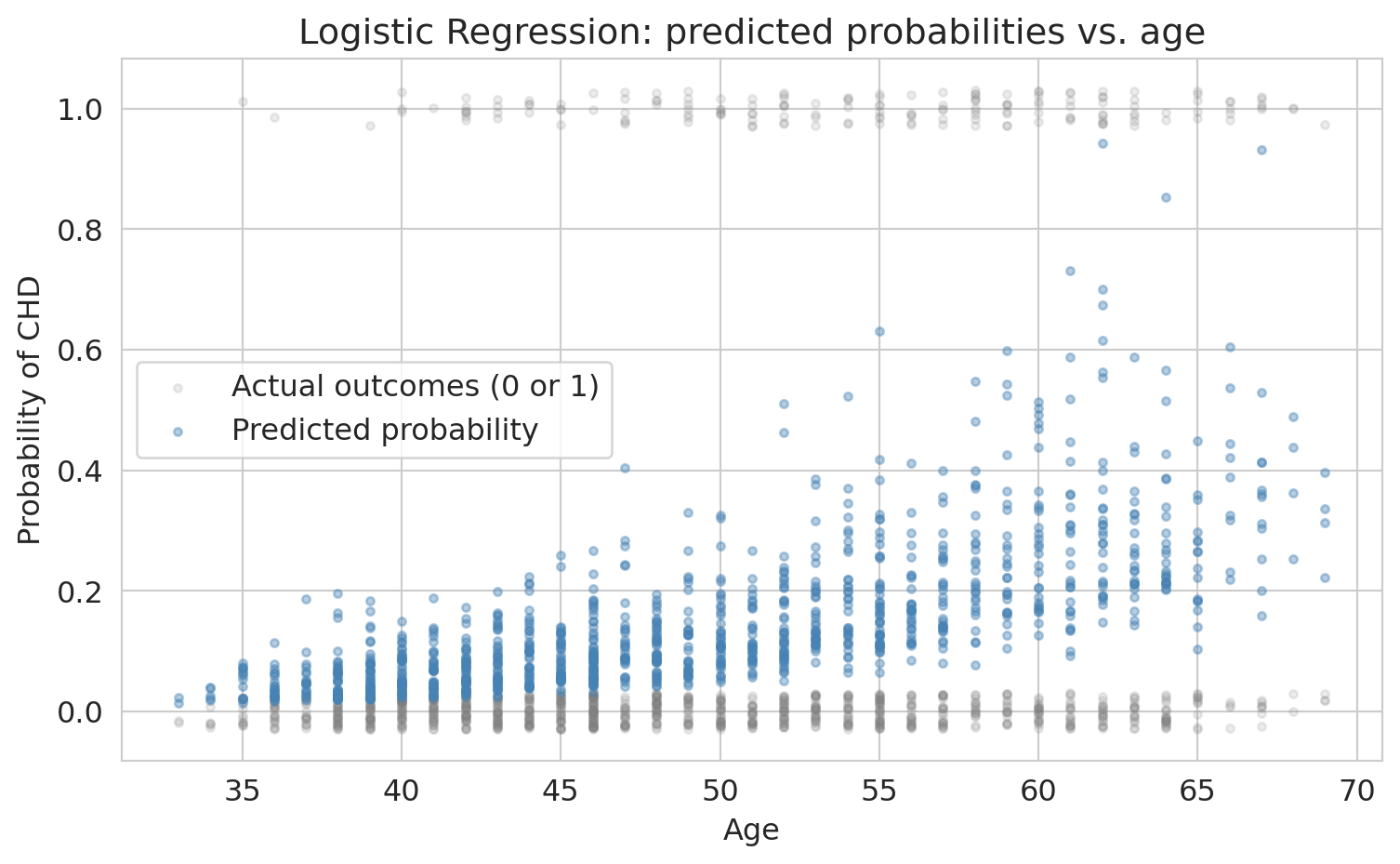
Let's see what the model's predictions look like on the data. Here we plot predicted CHD probability against age, with actual outcomes overlaid.
```{python}
# Visualize the logistic regression fit on data
y_proba_all = model.predict_proba(X_test_scaled)[:, 1]
fig, ax = plt.subplots(figsize=(8, 5))
# Jitter actual outcomes slightly for visibility
jitter = np.random.RandomState(42).uniform(-0.03, 0.03, size=len(y_test))
ax.scatter(X_test['age'], y_test + jitter, alpha=0.15, s=10,
color='gray', label='Actual outcomes (0 or 1)')
ax.scatter(X_test['age'], y_proba_all, alpha=0.4, s=10,
color='steelblue', label='Predicted probability')
ax.set_xlabel('Age')
ax.set_ylabel('Probability of CHD')
ax.set_title('Logistic Regression: predicted probabilities vs. age')
ax.legend()
plt.tight_layout()
plt.show()
```
The S-shape is visible: predicted probability increases with age, but stays between 0 and 1. Notice that the model's predicted probability is an *estimate* of each patient's true CHD risk (the *estimand*). Like all estimates, it comes with uncertainty.
Test accuracy of ~85%. Sounds great, right?
:::{.callout-tip}
## Think About It
Before we compare, what accuracy would you expect from a model that just predicts "no CHD" for every patient?
:::
**But wait.** What's the accuracy of the dumbest possible model — one that just predicts "no CHD" for everyone?
```{python}
# The "always predict negative" baseline
baseline_accuracy = 1 - y_test.mean()
print(f"Baseline accuracy (predict all negative): {baseline_accuracy:.3f}")
print(f"Our model accuracy: {model.score(X_test_scaled, y_test):.3f}")
print(f"Improvement over baseline: {model.score(X_test_scaled, y_test) - baseline_accuracy:.3f}")
```
Our fancy logistic regression barely beats "predict everyone is fine." That ~85% accuracy is **almost entirely driven by the class imbalance**, not by the model learning anything useful.
:::{.callout-warning}
## The Accuracy Trap
This phenomenon is the **accuracy trap** (also called the **accuracy paradox**). Accuracy measures how often you're right *overall*, but when one class dominates, being right about that class is easy and uninformative.
:::
## Interpreting coefficients: odds and odds ratios
Before we fix the evaluation problem, let's understand what the model learned. First, we need to understand **odds**.
If a patient's predicted CHD probability is $p = 0.20$, their **odds** are:
$$\text{odds} = \frac{p}{1-p} = \frac{0.20}{0.80} = 0.25 \quad \text{(or "1 to 4 against")}$$
If you've encountered odds in sports betting — "3 to 1 odds" — that's the same idea. Odds are just another way to express probability.
:::{.callout-important}
## Definition: Odds Ratio
The logit formula tells us that each coefficient $\beta_j$ affects the **log-odds** linearly. So $e^{\beta_j}$ is the **odds ratio** — the multiplicative change in odds per one-unit increase in $x_j$.
:::
Because features are standardized, each coefficient represents the change in log-odds per standard deviation increase, not per unit increase. If you fit without standardizing, the coefficient for age would tell you the odds change per one-year increase in age instead.
```{python}
# Create coefficient table
coef_df = pd.DataFrame({
'Feature': features,
'Coefficient': model.coef_[0],
'Odds Ratio': np.exp(model.coef_[0])
}).sort_values('Coefficient', ascending=False)
coef_df.round(3)
```
Each odds ratio tells us how the odds of CHD change per one-standard-deviation increase in that feature.
```{python}
# Interpret the coefficients
print("Logistic Regression Coefficients (standardized features):")
print("=" * 55)
for _, row in coef_df.iterrows():
direction = "+" if row['Coefficient'] > 0 else "-"
print(f" {row['Feature']:>12s}: odds x {row['Odds Ratio']:.2f} per 1-SD increase")
```
A concrete example: if a patient's predicted CHD probability is 0.20 (odds = 0.25), a one-SD increase in age multiplies those odds by about 1.7x, giving new odds of 0.424 — corresponding to a probability of about 30%.
The model says: age, systolic blood pressure, glucose, and male sex are the strongest risk factors. This ranking matches medical knowledge — but notice that the model treats each feature independently. It doesn't know that high blood pressure and high BMI often go together, or that young smokers face different risks than old smokers (for that, we'd need interaction terms — recall Chapter 6). We'll come back to this.
:::{.callout-tip}
## The Divide-by-4 Rule
Odds ratios are hard to feel. A quicker way to interpret logistic regression coefficients on the probability scale: **divide the coefficient by 4**. The result is the *maximum* possible change in $\Pr(Y=1)$ per one-unit increase in $x$.
Why does this work? The logistic curve is steepest at $p = 0.5$, where $dp/dx = p(1-p)\cdot\beta = 0.25\cdot\beta = \beta/4$. When $p$ is far from 0.5 the curve is flatter, so $\beta/4$ is an upper bound on the actual change.
**Example.** If the coefficient on "years of education" is 0.6, then each additional year of education increases the probability of the outcome by *at most* $0.6/4 = 0.15$ (15 percentage points). The true change is smaller when the baseline probability is near 0 or 1.
Source: Gelman & Hill, *Data Analysis Using Regression and Multilevel/Hierarchical Models* (2007).
:::
## What does the model predict for individual patients?
Before we evaluate overall performance, let's look at specific patients. This step makes the predicted probabilities concrete.
```{python}
# Pick a few patients to examine
y_proba = model.predict_proba(X_test_scaled)[:, 1]
test_examples = pd.DataFrame(X_test, columns=features)
test_examples['True CHD'] = y_test.values
test_examples['Predicted Prob'] = y_proba
# Show a few interesting cases
examples = test_examples.nsmallest(1, 'Predicted Prob')
examples = pd.concat([examples, test_examples.nlargest(2, 'Predicted Prob')])
# Find a borderline case near threshold 0.5
borderline = test_examples.iloc[(test_examples['Predicted Prob'] - 0.3).abs().argsort()[:1]]
examples = pd.concat([examples, borderline])
examples[['age', 'male', 'sysBP', 'cigsPerDay', 'True CHD', 'Predicted Prob']].round(3)
```
At the default threshold of 0.5, anyone with predicted probability above 0.5 is classified as "CHD." Is that the right threshold? We'll revisit this question soon.
## The confusion matrix: seeing where the model fails
:::{.callout-important}
## Definition: Confusion Matrix
The **confusion matrix** is a 2x2 table that cross-tabulates the model's predicted classes against the true classes, showing True Positives, False Positives, True Negatives, and False Negatives.
:::
Accuracy collapses everything into a single number. The confusion matrix shows us the full picture: what did the model get right, and *how* did it get things wrong?
```{python}
# Predictions at default threshold (0.5)
y_pred = model.predict(X_test_scaled)
# Confusion matrix
cm = confusion_matrix(y_test, y_pred)
fig, ax = plt.subplots(figsize=(6, 5))
disp = ConfusionMatrixDisplay(cm, display_labels=['No CHD', 'CHD'])
disp.plot(ax=ax, cmap='Blues', values_format='d')
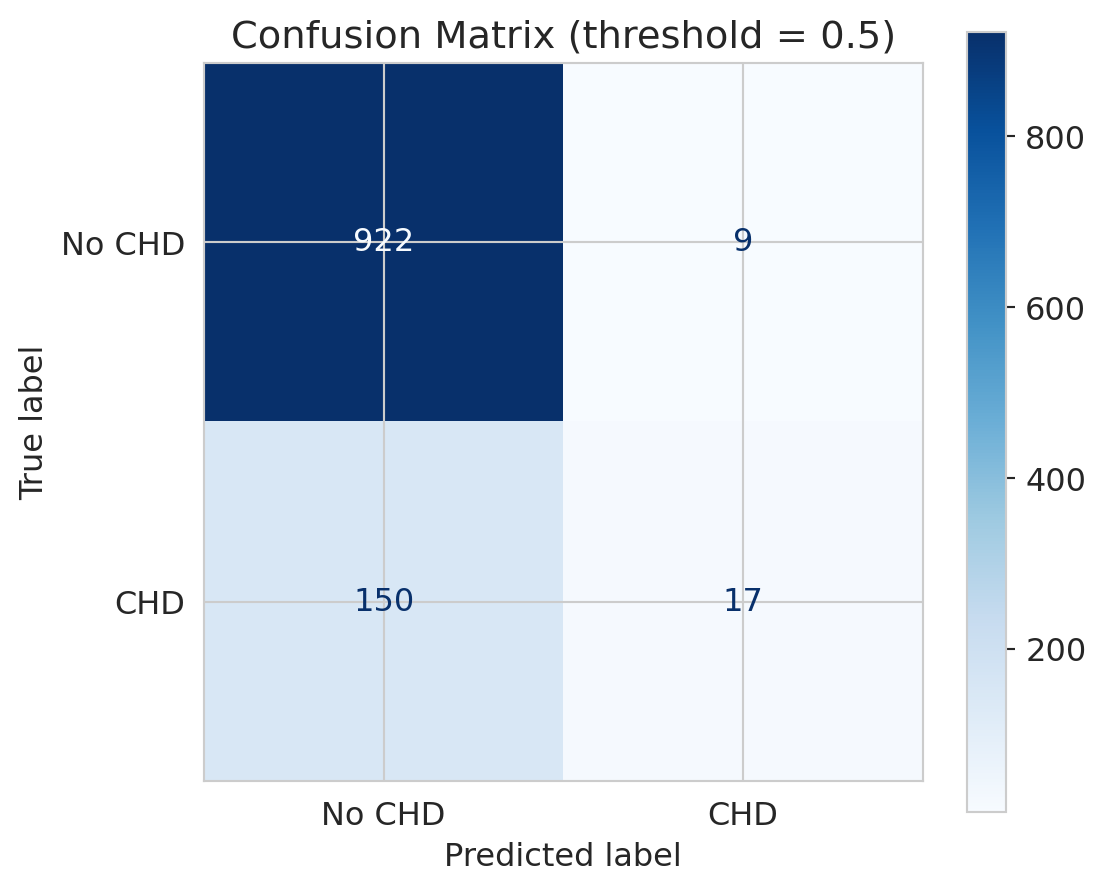
ax.set_title('Confusion Matrix (threshold = 0.5)')
plt.tight_layout()
plt.show()
# Print the numbers
tn, fp, fn, tp = cm.ravel()
print(f"True Negatives (correctly said 'no CHD'): {tn}")
print(f"False Positives (falsely alarmed): {fp}")
print(f"True Positives (correctly caught CHD): {tp}")
print(f"False Negatives (MISSED actual CHD): {fn}")
```
Look at the bottom row — actual CHD cases. The model catches very few of them! Most actual CHD patients are predicted as "no CHD." In a medical context, those are people who needed intervention and didn't get it.
This outcome is a **False Negative** — and in medicine, false negatives can be deadly.
:::{.callout-tip}
## Think About It
If you were designing a screening tool, which is worse — a false positive (unnecessary follow-up test) or a false negative (missed heart disease)?
:::
## Precision vs Recall
:::{.callout-important}
## Definition: Precision and Recall
- **Precision** = $\frac{TP}{TP + FP}$ = "Of those I flagged as CHD, how many really had it?"
- **Recall** (Sensitivity) = $\frac{TP}{TP + FN}$ = "Of all actual CHD cases, how many did I catch?"
:::
A mnemonic: Precision asks, "I flagged this patient — should I trust my flag?" Recall asks, "Of all sick people, how many did we find?"
Two metrics capture the two types of mistakes, and they tell very different stories.
```{python}
# Precision and recall at default threshold
precision = tp / (tp + fp) if (tp + fp) > 0 else 0
recall = tp / (tp + fn)
print(f"Precision: {precision:.3f}")
print(f" → Of patients flagged as CHD, {precision*100:.0f}% actually had it")
print()
print(f"Recall: {recall:.3f}")
print(f" → Of actual CHD patients, we caught only {recall*100:.0f}%")
print()
print(f"Accuracy: {(tp+tn)/(tp+tn+fp+fn):.3f}")
print(f" → This number hides the fact that recall is terrible")
```
Precision is decent — when the model *does* flag someone, it's often right. But recall is abysmal: we're missing the vast majority of actual CHD cases. The low base rate makes the model **too conservative** — it rarely predicts CHD.
Note: if the model never predicts positive, precision is undefined (0/0). We set it to 0 by convention, but the edge case is worth noting — with such a conservative model, it nearly occurs here.
The **F1 score** is the harmonic mean of precision and recall: $F_1 = 2 \cdot \frac{\text{precision} \cdot \text{recall}}{\text{precision} + \text{recall}}$. It balances both concerns in a single number, but it still doesn't capture the *cost* of each type of mistake.
This tension is the fundamental tradeoff: catching more positives (higher recall) means accepting more false alarms (lower precision).
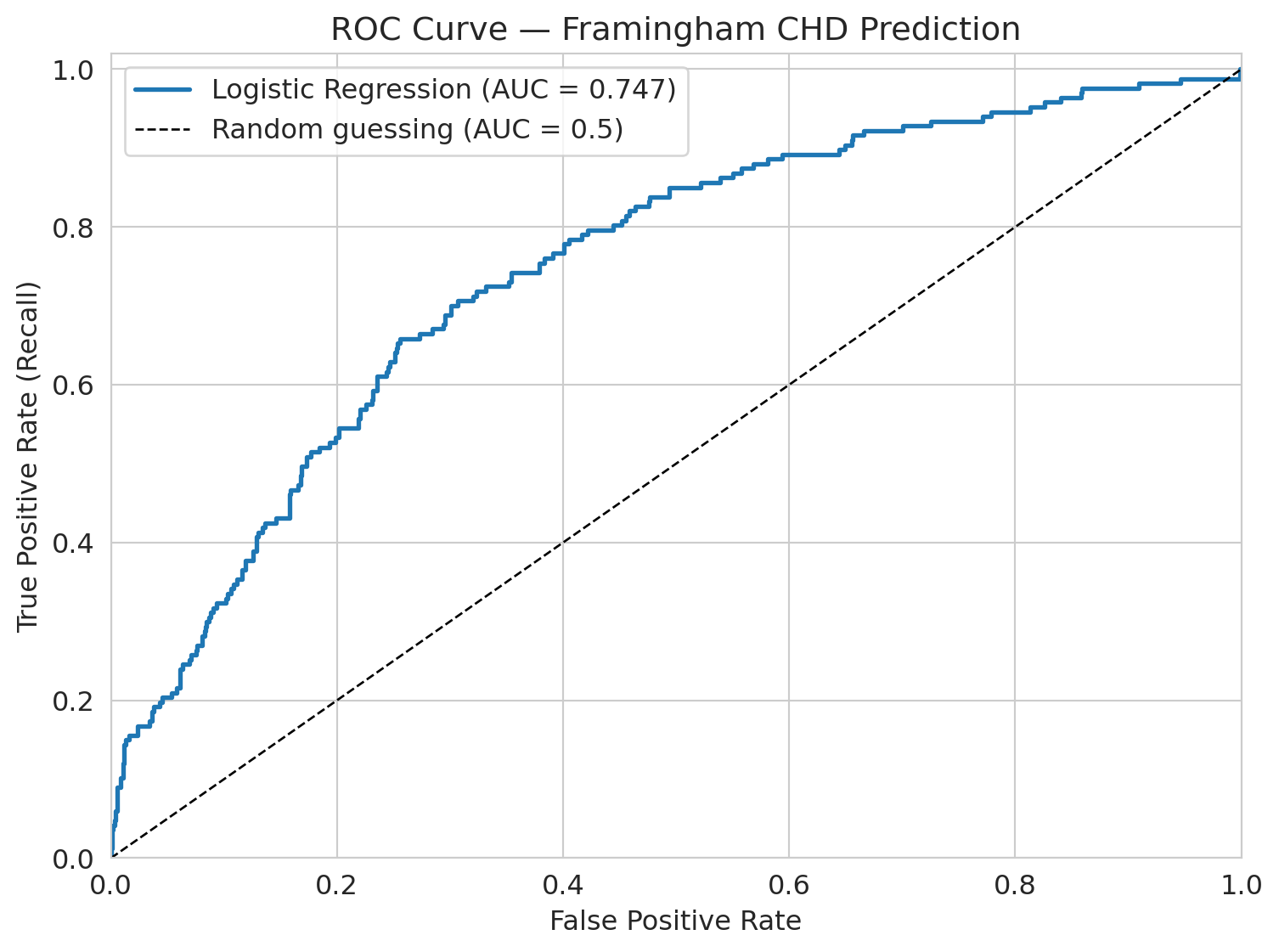
## The ROC curve: evaluating across all thresholds
So far we've been using the default threshold of 0.5: if the predicted probability is above 0.5, predict CHD. But why 0.5?
:::{.callout-important}
## Definition: ROC Curve and AUC
The **ROC curve** shows model performance across *all* possible thresholds. It plots the True Positive Rate (recall) against the False Positive Rate at each threshold.
The **AUC** (area under the ROC curve) summarizes this: a perfect model has AUC = 1.0, and random guessing gives AUC = 0.5.
:::
```{python}
# Get predicted probabilities
y_proba = model.predict_proba(X_test_scaled)[:, 1]
# ROC curve
fpr, tpr, thresholds_roc = roc_curve(y_test, y_proba)
roc_auc = auc(fpr, tpr)
fig, ax = plt.subplots(figsize=(8, 6))
ax.plot(fpr, tpr, linewidth=2, label=f'Logistic Regression (AUC = {roc_auc:.3f})')
ax.plot([0, 1], [0, 1], 'k--', linewidth=1, label='Random guessing (AUC = 0.5)')
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate (Recall)')
ax.set_title('ROC Curve — Framingham CHD Prediction')
ax.legend(fontsize=12)
ax.set_xlim([0, 1])
ax.set_ylim([0, 1.02])
plt.tight_layout()
plt.show()
print(f"AUC = {roc_auc:.3f}")
print("Interpretation: if you pick one CHD patient and one non-CHD patient")
print(f"at random, the model ranks the CHD patient higher {roc_auc*100:.0f}% of the time.")
print()
print("This quantity is the concordance probability. It measures the model's")
print("ability to *rank* patients by risk -- not how well it classifies at any threshold.")
```
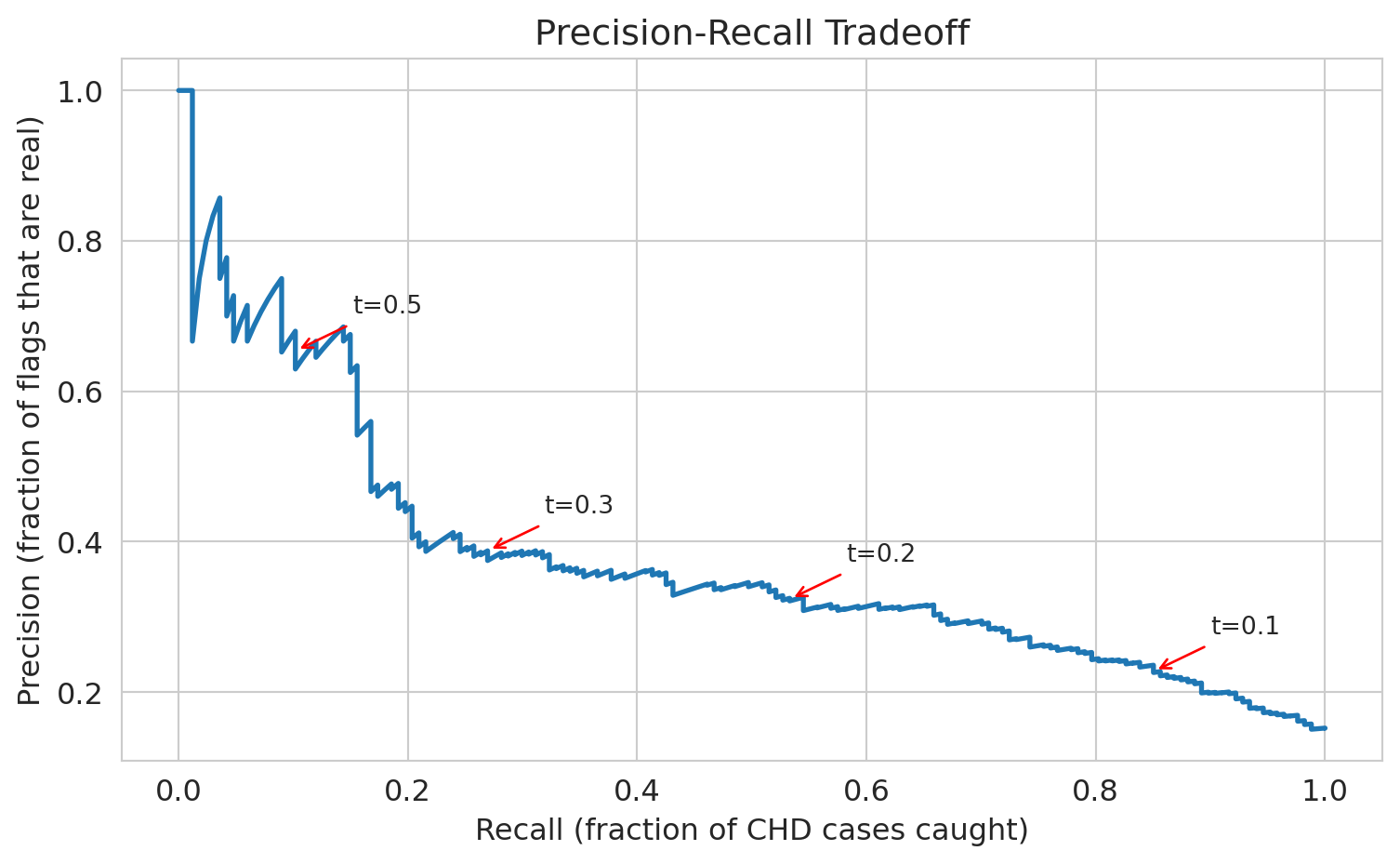
## Threshold choice: the precision-recall tradeoff
The default threshold of 0.5 gives us terrible recall. What if we lower it?
If we set the threshold to 0.3 — flag anyone with >30% predicted CHD risk — we'll catch more true cases but also flag more false positives. Let's see the tradeoff.
```{python}
# Compare thresholds
thresholds_to_try = [0.5, 0.4, 0.3, 0.2, 0.1]
print(f"{'Threshold':>10s} {'Precision':>10s} {'Recall':>10s} {'Accuracy':>10s} {'Flagged':>10s}")
print("=" * 55)
# For each threshold, classify patients and compute metrics
for t in thresholds_to_try:
y_pred_t = (y_proba >= t).astype(int)
cm_t = confusion_matrix(y_test, y_pred_t)
tn_t, fp_t, fn_t, tp_t = cm_t.ravel()
prec_t = tp_t / (tp_t + fp_t) if (tp_t + fp_t) > 0 else 0
rec_t = tp_t / (tp_t + fn_t)
acc_t = (tp_t + tn_t) / (tp_t + tn_t + fp_t + fn_t)
flagged = tp_t + fp_t
print(f"{t:>10.2f} {prec_t:>10.3f} {rec_t:>10.3f} {acc_t:>10.3f} {flagged:>10d}")
```
See the tradeoff? As we lower the threshold:
- **Recall goes up** — we catch more CHD cases
- **Precision goes down** — more false alarms
- **Accuracy goes down** — more negative cases are now misclassified
There's no free lunch. The right threshold depends on the **cost of each mistake** — a consequential decision, not a statistical one.
If a follow-up test costs \$500 and a missed CHD case costs \$50,000 in emergency care, what threshold minimizes expected cost? In medicine, missing a heart disease case is far worse than ordering an extra test, so we'd choose a low threshold. In spam filtering, false positives (real email in spam) are worse, so we'd choose a high threshold.
:::{.callout-tip}
## Think About It
If you're the hospital administrator and you have budget for follow-up appointments with 200 patients per month, which threshold would you choose? This choice isn't a statistics question — it's a resource allocation question.
:::
```{python}
# Precision-Recall curve
precision_curve, recall_curve, thresholds_pr = precision_recall_curve(y_test, y_proba)
fig, ax = plt.subplots(figsize=(8, 5))
ax.plot(recall_curve, precision_curve, linewidth=2)
ax.set_xlabel('Recall (fraction of CHD cases caught)')
ax.set_ylabel('Precision (fraction of flags that are real)')
ax.set_title('Precision-Recall Tradeoff')
# Annotate a few thresholds
for t in [0.1, 0.2, 0.3, 0.5]:
# Find the index of the threshold closest to t
idx = np.argmin(np.abs(thresholds_pr - t))
ax.annotate(f't={t}', xy=(recall_curve[idx], precision_curve[idx]),
fontsize=10, ha='center',
arrowprops=dict(arrowstyle='->', color='red'),
xytext=(recall_curve[idx]+0.08, precision_curve[idx]+0.05))
plt.tight_layout()
plt.show()
```
## Who does the model fail for?
The overall AUC is ~0.75. Not amazing, but not terrible. But *overall* performance can hide dramatic failures in subgroups.
:::{.callout-tip}
## Think About It
Before we look at the numbers — for which age group do you think the model performs worst? Why?
:::
Let's check: does the model work equally well for all ages?
```{python}
# Create age groups in the test set
test_df = pd.DataFrame(X_test, columns=features)
test_df['y_true'] = y_test.values
test_df['y_proba'] = y_proba
# Decade-based age groups for interpretability
test_df['age_group'] = pd.cut(test_df['age'], bins=[30, 40, 50, 60, 70],
labels=['30-40', '40-50', '50-60', '60-70'])
# AUC by age group
print(f"{'Age Group':>10s} {'N':>6s} {'CHD Rate':>10s} {'AUC':>8s}")
print("=" * 40)
for group in ['30-40', '40-50', '50-60', '60-70']:
mask = test_df['age_group'] == group
sub = test_df[mask]
n = len(sub)
chd_rate = sub['y_true'].mean()
if sub['y_true'].nunique() < 2:
auc_val = float('nan')
else:
fpr_g, tpr_g, _ = roc_curve(sub['y_true'], sub['y_proba'])
auc_val = auc(fpr_g, tpr_g)
print(f"{group:>10s} {n:>6d} {chd_rate:>10.3f} {auc_val:>8.3f}")
```
**There it is.** For patients under 40, the AUC drops dramatically — close to 0.5, which is *random guessing*. The model essentially learned "older people get heart disease" and has almost no ability to identify risk in younger patients.
This failure is critical for a screening tool. Young patients who *do* develop CHD are exactly the cases where early intervention matters most — and the model is blind to them.
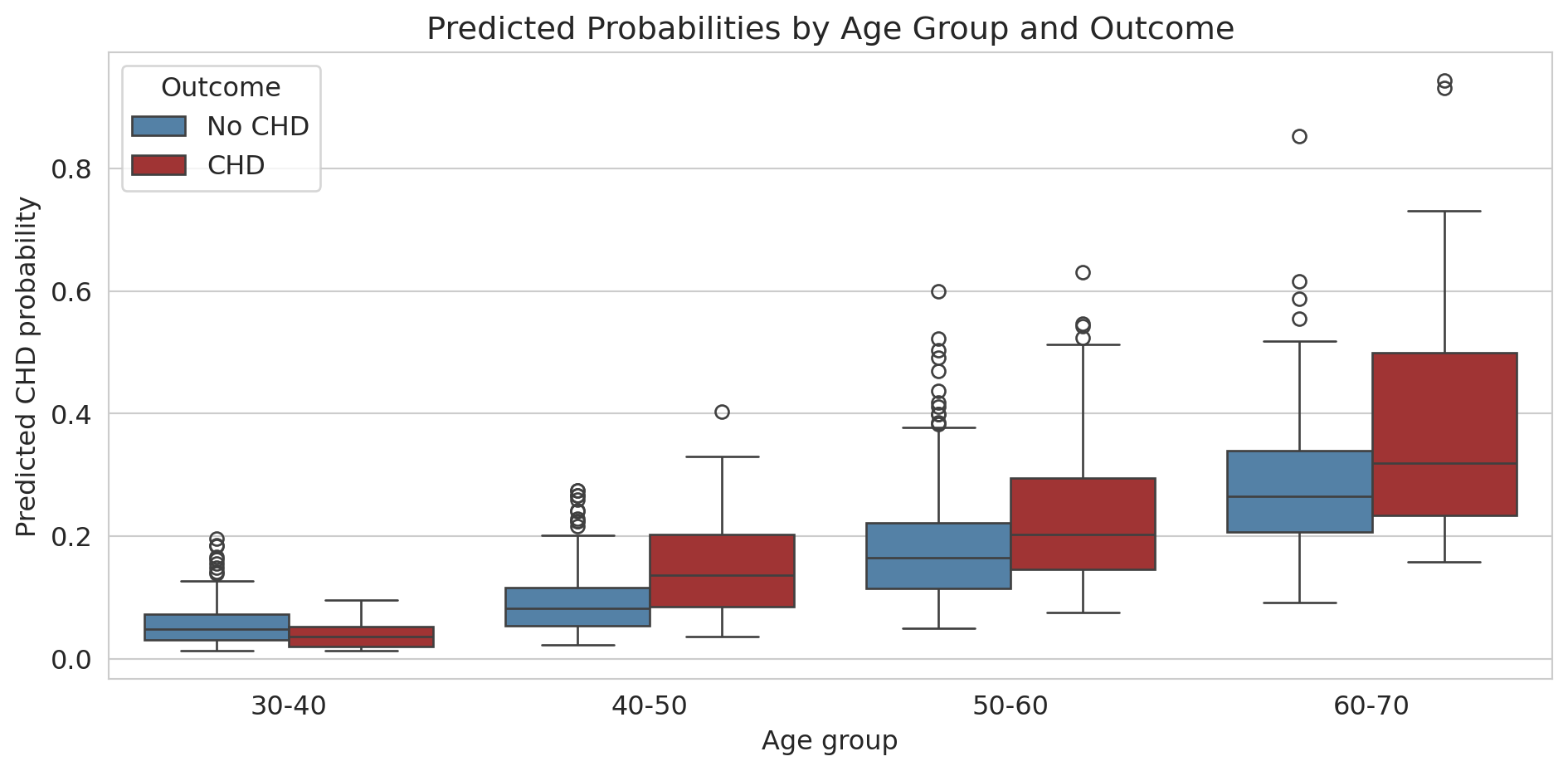
```{python}
# Box plot: predicted probabilities by age group and actual outcome
test_df['Outcome'] = test_df['y_true'].map({0: 'No CHD', 1: 'CHD'})
fig, ax = plt.subplots(figsize=(10, 5))
sns.boxplot(data=test_df.dropna(subset=['age_group']),
x='age_group', y='y_proba', hue='Outcome',
palette={'No CHD': 'steelblue', 'CHD': 'firebrick'}, ax=ax)
ax.set_xlabel('Age group')
ax.set_ylabel('Predicted CHD probability')
ax.set_title('Predicted Probabilities by Age Group and Outcome')
plt.tight_layout()
plt.show()
```
For older patients, the model separates CHD from non-CHD reasonably well (the distributions don't overlap as much). For younger patients, the predicted probabilities are all bunched near zero — the model thinks *nobody* young is at risk.
**Why this matters:** An AI assistant would report the overall AUC of 0.75, declare the model "moderately good," and move on. It wouldn't check subgroup performance. But in practice, a model that fails for young patients is dangerous — it creates a false sense of security for exactly the people who most need screening.
What could we do about it? Options include training separate models for different age groups, adding interaction terms (recall Chapter 6), or collecting more data on young CHD patients. There's no easy fix — but the first step is always checking.
## Same problem, different domain: hospital readmissions
The Framingham CHD prediction and hospital readmission prediction share the same fundamental challenge: class imbalance. In the CMS hospital readmissions data, about 15-20% of patients are readmitted within 30 days — almost the same positive rate as our CHD data. The same traps apply: accuracy is misleading, the "always predict negative" model looks great on paper, and threshold choice depends on the cost of each type of mistake.
The hospital has to decide: **where on the precision-recall curve do we want to operate?** There's no purely statistical answer — it depends on intervention costs, staff availability, and patient outcomes.
## Key Takeaways
- **Logistic regression** predicts probabilities using the sigmoid function. The **logit** (log-odds) is a linear function of the features, and coefficients tell you the multiplicative change in odds.
- **The loss function determines what the model finds.** Linear regression minimizes squared error. Logistic regression minimizes logistic loss (cross-entropy). The choice of loss function is a modeling decision.
- **Gradient descent** finds the coefficients by iteratively walking downhill on the loss landscape. For logistic regression (convex), it finds THE answer. For neural networks (non-convex), you might get stuck in a local minimum.
- **Accuracy is misleading** with imbalanced classes. A model that predicts "everyone is fine" gets 85% accuracy but catches zero actual cases.
- **The confusion matrix** reveals what accuracy hides: look at True Positives, False Positives, True Negatives, and False Negatives separately.
- **Precision vs Recall** captures the fundamental tradeoff: catching more positives (recall) means more false alarms (lower precision). The right balance depends on the cost of each type of mistake — a consequential decision.
- **Always check subgroup performance.** An overall AUC of 0.75 can hide the fact that the model is *useless* for certain patient groups. Models that work "on average" can fail exactly where it matters most.
## Metrics cheat sheet
| Metric | Formula | What it measures |
|--------|---------|-----------------|
| **Accuracy** | (TP + TN) / N | Overall correctness (misleading with imbalance) |
| **Precision** | TP / (TP + FP) | Of those flagged, how many are real? |
| **Recall** | TP / (TP + FN) | Of real cases, how many caught? |
| **F1 score** | 2 * Prec * Rec / (Prec + Rec) | Harmonic mean of precision and recall |
| **AUC** | Area under ROC curve | Overall ranking ability (threshold-free) |
## Study guide
### Key ideas
1. Logistic regression models the probability of a binary outcome by passing a linear combination through the sigmoid function.
2. OLS has a closed-form solution; logistic regression does not. The sigmoid makes the loss nonlinear, requiring gradient descent.
3. The logistic loss is the negative log-likelihood under a Bernoulli model — the Bernoulli distribution models a single binary outcome with probability $p$, and logistic regression estimates that $p$ as a function of features. This loss penalizes confident wrong predictions heavily.
4. Gradient descent finds the coefficients by iteratively stepping in the direction that reduces the loss; convexity guarantees convergence to the global minimum.
5. Accuracy is misleading with class imbalance — always look at precision, recall, and the confusion matrix.
6. Threshold choice is a cost-benefit decision, not a statistical one.
7. Subgroup analysis can reveal dramatic failures hidden by aggregate performance.
### Definitions
- **Sigmoid function**: maps any real number to $[0, 1]$, converting linear predictions to probabilities
- **Logit (log-odds)**: $\log(p/(1-p))$, the inverse of the sigmoid; linear in the features
- **Odds ratio**: $e^{\beta_j}$, the multiplicative change in odds per unit increase in $x_j$
- **Logistic loss (cross-entropy)**: the loss function logistic regression minimizes; heavily penalizes confident wrong predictions
- **Gradient descent**: iterative optimization algorithm that follows the negative gradient downhill
- **Learning rate**: step size in gradient descent; too large causes overshoot, too small causes slow convergence
- **Convex**: a loss landscape with a single valley; gradient descent finds the global minimum
- **Confusion matrix**: 2x2 table of TP, FP, TN, FN
- **Precision**: fraction of positive predictions that are correct
- **Recall (sensitivity)**: fraction of actual positives that are caught
- **F1 score**: harmonic mean of precision and recall
- **ROC curve**: True Positive Rate vs. False Positive Rate across all thresholds
- **AUC**: area under the ROC curve; 1.0 = perfect, 0.5 = random guessing
### Computational tools
- `LogisticRegression()` — fit a logistic regression model
- `model.predict_proba()` — get predicted probabilities (not just 0/1 predictions)
- `confusion_matrix()` — compute the 2x2 confusion matrix
- `ConfusionMatrixDisplay()` — plot the confusion matrix
- `roc_curve()`, `auc()` — compute the ROC curve and its area
- `precision_recall_curve()` — compute the precision-recall curve
- `classification_report()` — print precision, recall, F1, and accuracy in one call
### For the quiz
Be able to (1) interpret logistic regression coefficients as odds ratios, (2) compute precision and recall from a confusion matrix, (3) explain why accuracy is misleading with class imbalance, (4) describe the gradient descent update rule and what convexity guarantees, (5) choose a threshold given the costs of false positives and false negatives.
## Coming up next
Act 3 continues with [Chapter 14](lec14-pca.qmd) on **PCA** — when you have 96 columns, can you summarize them with just 5? PCA finds the best low-dimensional summary of your data, and the linear algebra from earlier in the course tells us exactly what "best" means.
And in **Chapter 17**, we'll see how gradient descent powers **gradient boosting** — an algorithm that builds on today's ideas to produce state-of-the-art predictions.